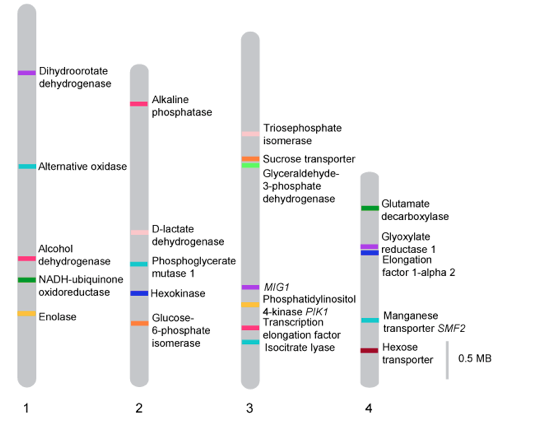
The strain Dekkera bruxellensis CBS 11270 has been isolated from a Swedish alcohol production plant, where it outcompeted the initial inoculum of Saccharomyces cerevisiae. In order to understand genome evolution of a D. bruxellensis strain, which was exposed to conditions of years-long continuous industrial fermentation, we analyzed it using next generation sequencing technologies (NGS). We developed an improved efficient approach for de-novo assembly of genomes of non-conventional yeasts, enabling us assembly of scaffolds in chromosome size. This method is based on multi technology approach combining short and long-read sequencing and Opgen optical mapping. The genome of D. bruxellensis CBS 11270 was shown to be diploid and composed of four chromosomes. Chromosomal genome assembly allowed identifying several inter- and intrachromosomal genes duplications. When the scaffolds of the CBS 2499 genome assembly were aligned against the chromosomes of CBS 11270, some scaffolds were split and aligned in different order or to different chromosomes. These results indicated adaptation of CBS 11270 to stable conditions of industrial fermentation and indicates a high genome flexibility of the yeast species D. bruxellensis.
Publication:
De novo assembly of Dekkera bruxellensis: a multi technology approach using short and long-read sequencing and optical mapping. Remi-Andre Olsen, Ignas Bunikis, Tiukova I., Holmberg K., Britta Lötstedt B., Vinnere Pettersson O., Passoth V., Käller M., Vezzi F., Lotstedt B., Kaller M. (2015) GIGASCIENCE 4